GLUT1

| glucose transporter, type 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | Glu_transpt_1IPR002439erythrocyte/brain hexose facilitatorGLUT1glucose transporter-1Gtr1Glut1Glut-1Glucose Transporter Type 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | GeneCards: [1]; OMA:- orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Glucose transporter 1 (or GLUT1), also known as solute carrier family 2, facilitated glucose transporter member 1 (SLC2A1), is a uniporter protein that in humans is encoded by the SLC2A1 gene.[1] GLUT1 facilitates the transport of glucose across the plasma membranes of mammalian cells.[2] This gene encodes a facilitative glucose transporter that is highly expressed in erythrocytes and endothelial cells, including cells of the blood–brain barrier. The encoded protein is found primarily in the cell membrane and on the cell surface, where it can also function as a receptor for human T-cell leukemia virus (HTLV) I and II.[3] GLUT1 accounts for 2 percent of the protein in the plasma membrane of erythrocytes.
Mutations in this gene can cause GLUT1 deficiency syndrome 1, GLUT1 deficiency syndrome 2, idiopathic generalized epilepsy 12, dystonia 9, and stomatin-deficient cryohydrocytosis.[4][5]
Discovery
[edit]GLUT1 was the first glucose transporter to be characterized. GLUT 1 is highly conserved.[1] GLUT 1 of humans and mice have 98% identity at the amino acid level. GLUT 1 is encoded by the SLC2 gene and is one of a family of 14 genes encoding GLUT proteins.[6]
Structure
[edit]The SLC2A1 gene is located on the p arm of chromosome 1 in position 34.2 and has 10 exons spanning 33,802 base pairs.[3] The gene produces a 54.1 kDa protein composed of 492 amino acids.[7][8][9][10] It is a multi-pass protein located in the cell membrane.[4][5] This protein lacks a signal sequence; its C-terminus, N-terminus, and the very hydrophilic domain in the protein's center are all predicted to lie on the cytoplasmic side of the cell membrane.[10][1]
GLUT1 behaves as a Michaelis–Menten enzyme and contains 12 membrane-spanning alpha helices, each containing 20 amino acid residues. A helical wheel analysis shows that the membrane-spanning alpha-helices are amphipathic, with one side being polar and the other side hydrophobic. Six of these membrane-spanning helices are believed to bind together in the membrane to create a polar channel in the center through which glucose can traverse, with the hydrophobic regions on the outside of the channel adjacent to the fatty acid tails of the membrane.[citation needed]
Function
[edit]Energy-yielding metabolism in erythrocytes depends on a constant supply of glucose from the blood plasma, where the glucose concentration is maintained at about 5mM. Glucose enters the erythrocyte by facilitated diffusion via a specific glucose transporter, at a rate of about 50,000 times greater than uncatalyzed transmembrane diffusion. The glucose transporter of erythrocytes (called GLUT1 to distinguish it from related glucose transporters in other tissues) is a type III integral protein with 12 hydrophobic segments, each of which is believed to form a membrane-spanning helix. The detailed structure of GLUT1 is not known yet, but one plausible model suggests that the side-by-side assembly of several helices produces a transmembrane channel lined with hydrophilic residues that can hydrogen-bond with glucose as it moves through the channel.[11]
GLUT1 is responsible for the low level of basal glucose uptake required to sustain respiration in all cells. Expression levels of GLUT1 in cell membranes are increased by reduced glucose levels and decreased by increased glucose levels.[citation needed]
GLUT1 is also a major receptor for uptake of Vitamin C as well as glucose, especially in non vitamin C producing mammals as part of an adaptation to compensate by participating in a Vitamin C recycling process. In mammals that do produce Vitamin C, GLUT4 is often expressed instead of GLUT1.[12]
Tissue distribution
[edit]GLUT1 expression occurs in almost all tissues, with the degree of expression typically correlating with the rate of cellular glucose metabolism. In the adult it is expressed at highest levels in erythrocytes and also in the endothelial cells of barrier tissues such as the blood–brain barrier.[13]
Clinical significance
[edit]Mutations in the GLUT1 gene are responsible for GLUT1 deficiency or De Vivo disease, which is a rare autosomal dominant disorder.[14] This disease is characterized by a low cerebrospinal fluid glucose concentration (hypoglycorrhachia), a type of neuroglycopenia, which results from impaired glucose transport across the blood–brain barrier.
GLUT1 Deficiency Syndrome 1
[edit]Many mutations in the SLC2A1 gene, including LYS456TER, TYR449TER, LYS256VAL, ARG126HIS, ARG126LEU and GLY91ASP, have been shown to cause GLUT1 deficiency syndrome 1 (GLUT1DS1), a neurologic disorder showing wide phenotypic variability. This disease can be inherited in either an autosomal recessive or autosomal dominant manner.[10] The most severe 'classic' phenotype comprises infantile-onset epileptic encephalopathy associated with delayed development, acquired microcephaly, motor incoordination, and spasticity. Onset of seizures, usually characterized by apneic episodes, staring spells, and episodic eye movements, occurs within the first 4 months of life. Other paroxysmal findings include intermittent ataxia, confusion, lethargy, sleep disturbance, and headache. Varying degrees of cognitive impairment can occur, ranging from learning disabilities to severe mental retardation.[4][5]
GLUT1 Deficiency Syndrome 2
[edit]Other mutations, like GLY314SER, ALA275THR, ASN34ILE, SER95ILE, ARG93TRP, ARG91TRP, a 3-bp insertion (TYR292) and a 12-bp deletion (1022_1033del) in exon 6, have been shown to cause GLUT1 deficiency syndrome 2 (GLUT1DS2), a clinically variable disorder characterized primarily by onset in childhood of paroxysmal exercise-induced dyskinesia. The dyskinesia involves transient abnormal involuntary movements, such as dystonia and choreoathetosis, induced by exercise or exertion, and affecting the exercised limbs. Some patients may also have epilepsy, most commonly childhood absence epilepsy. Mild mental retardation may also occur. In some patients involuntary exertion-induced dystonic, choreoathetotic, and ballistic movements may be associated with macrocytic hemolytic anemia.[4][5] Inheritance of this disease is autosomal dominant.[10]
Idiopathic Generalized Epilepsy 12
[edit]Some mutations, particularly ASN411SER, ARG458TRP, ARG223PRO and ARG232CYS, have been shown to cause idiopathic generalized epilepsy 12 (EIG12), a disorder characterized by recurring generalized seizures in the absence of detectable brain lesions and/or metabolic abnormalities. Generalized seizures arise diffusely and simultaneously from both hemispheres of the brain. Seizure types include juvenile myoclonic seizures, absence seizures, and generalized tonic-clonic seizures. In some EIG12 patients seizures may remit with age.[4][5] Inheritance of this disease is autosomal dominant.[10]
Dystonia 9
[edit]Another mutation, ARG212CYS, has been shown to cause Dystonia 9 (DYT9), an autosomal dominant neurologic disorder characterized by childhood onset of paroxysmal choreoathetosis and progressive spastic paraplegia. Most patients show some degree of cognitive impairment. Other variable features may include seizures, migraine headaches, and ataxia.[4][5]
Stomatin-deficient Cryohydrocytosis
[edit]Certain mutations, like GLY286ASP and a 3-bp deletion in ILE435/436, cause Stomatin-deficient cryohydrocytosis with neurologic defects (SDCHCN), a rare form of stomatocytosis characterized by episodic hemolytic anemia, cold-induced red cells cation leak, erratic hyperkalemia, neonatal hyperbilirubinemia, hepatosplenomegaly, cataracts, seizures, mental retardation, and movement disorder.[4][5] Inheritance of this disease is autosomal dominant.[10]
Role as a Receptor for HTLV
[edit]GLUT1 is also a receptor used by the HTLV virus to gain entry into target cells.[15]
Role as a Histochemical Marker for Hemangioma
[edit]Glut1 has also been demonstrated as a powerful histochemical marker for hemangioma of infancy[16]
Interactions
[edit]GLUT1 has been shown to interact with GIPC1.[17] It is found in a complex with Adducin (ADD2) and Dematin (EPB49) and interacts (via C-terminus cytoplasmic region) with Dematin isoform 2.[18] It also interacts with SNX27; the interaction is required when endocytosed to prevent degradation in lysosomes and promote recycling to the plasma membrane.[19] This protein interacts with STOM.[20] It interacts with SGTA (via Gln-rich region) and has binary interactions with CREB3-2.[4][5]
GLUT1 has two significant types in the brain: 45-kDa and 55-kDa. GLUT1 45-kDa is present in astroglia and neurons. GLUT1 55-kDa is present in the endothelial cells of the brain vasculature and is responsible for glucose transport across the blood–brain barrier; its deficiency causes a low level of glucose in CSF (less than 60 mg/dl) which may elicit seizures in deficient individuals.[citation needed]
Recently a GLUT1 inhibitor DERL3 has been described and is often methylated in colorectal cancer. In this cancer, DERL3 methylations seem to mediate the Warburg effect.[21]
Inhibitors
[edit]Fasentin is a small molecule inhibitor of the intracellular domain of GLUT1 preventing glucose uptake.[22]
Recently, a new more selective GLUT1 inhibitor, Bay-876, has been described.[23]
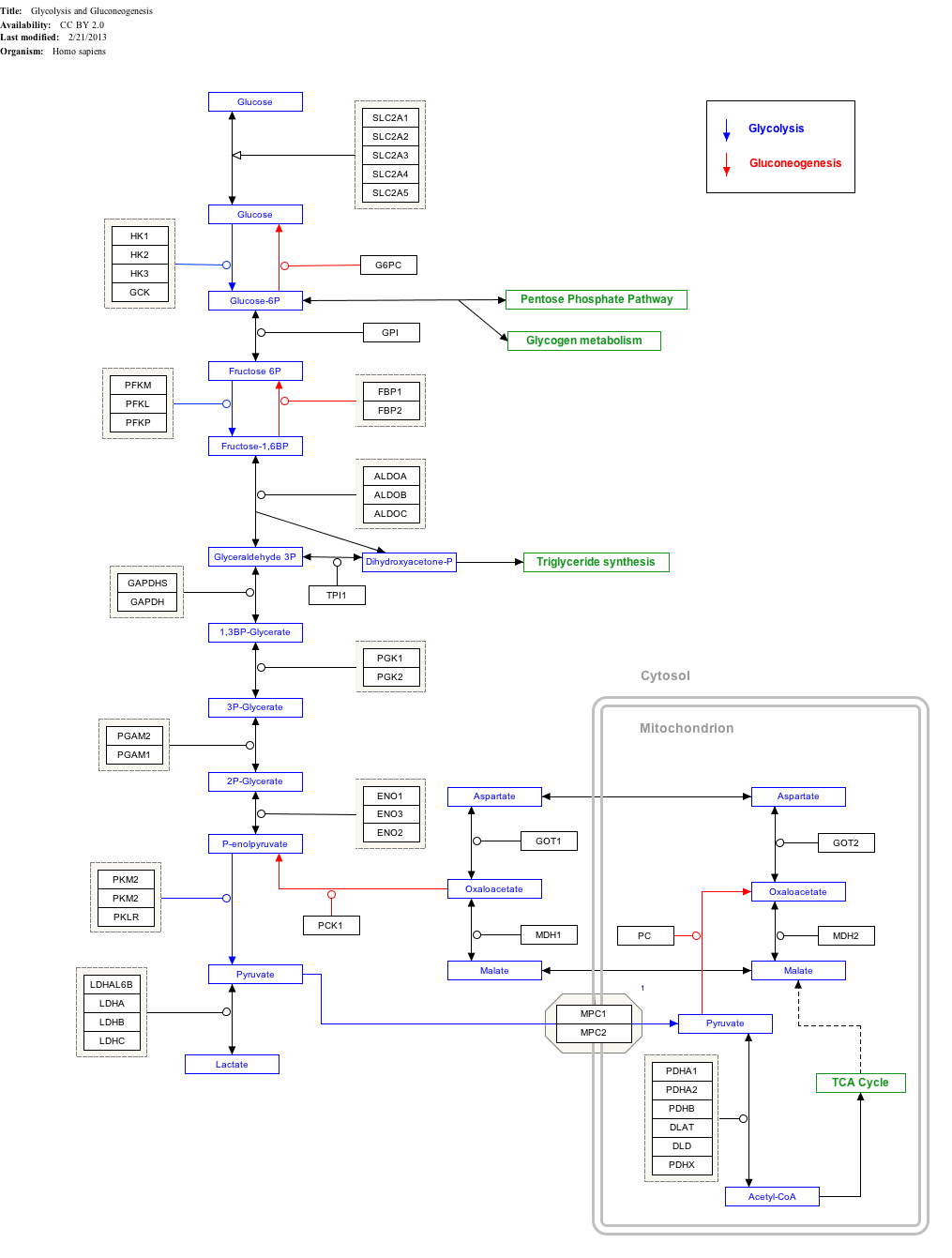
Interactive pathway map
[edit]Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "GlycolysisGluconeogenesis_WP534".
References
[edit]- ^ a b c Mueckler M, Caruso C, Baldwin SA, Panico M, Blench I, Morris HR, Allard WJ, Lienhard GE, Lodish HF (September 1985). "Sequence and structure of a human glucose transporter". Science. 229 (4717): 941–5. Bibcode:1985Sci...229..941M. doi:10.1126/science.3839598. PMID 3839598.
- ^ Olson AL, Pessin JE (1996). "Structure, function, and regulation of the mammalian facilitative glucose transporter gene family". Annual Review of Nutrition. 16: 235–56. doi:10.1146/annurev.nu.16.070196.001315. PMID 8839927.
- ^ a b
 This article incorporates text from this source, which is in the public domain: "Entrez Gene: Transmembrane protein 70". Retrieved 2018-08-14.
This article incorporates text from this source, which is in the public domain: "Entrez Gene: Transmembrane protein 70". Retrieved 2018-08-14. - ^ a b c d e f g h "SLC2A1 – Solute carrier family 2, facilitated glucose transporter member 1 – Homo sapiens (Human) – SLC2A1 gene & protein". www.uniprot.org. Retrieved 2018-08-27.
 This article incorporates text available under the CC BY 4.0 license.
This article incorporates text available under the CC BY 4.0 license. - ^ a b c d e f g h "UniProt: the universal protein knowledgebase". Nucleic Acids Research. 45 (D1): D158–D169. January 2017. doi:10.1093/nar/gkw1099. PMC 5210571. PMID 27899622.
- ^ Mueckler M, Thorens B (2013). "The SLC2 (GLUT) family of membrane transporters". Molecular Aspects of Medicine. 34 (2–3): 121–38. doi:10.1016/j.mam.2012.07.001. PMC 4104978. PMID 23506862.
- ^ Zong NC, Li H, Li H, Lam MP, Jimenez RC, Kim CS, Deng N, Kim AK, Choi JH, Zelaya I, Liem D, Meyer D, Odeberg J, Fang C, Lu HJ, Xu T, Weiss J, Duan H, Uhlen M, Yates JR, Apweiler R, Ge J, Hermjakob H, Ping P (October 2013). "Integration of cardiac proteome biology and medicine by a specialized knowledgebase". Circulation Research. 113 (9): 1043–53. doi:10.1161/CIRCRESAHA.113.301151. PMC 4076475. PMID 23965338.
- ^ "SLC2A1 – Solute carrier family 2, facilitated glucose transporter member 1". Cardiac Organellar Protein Atlas Knowledgebase (COPaKB).
- ^ Wang D, Kranz-Eble P, De Vivo DC (September 2000). "Mutational analysis of GLUT1 (SLC2A1) in Glut-1 deficiency syndrome". Human Mutation. 16 (3): 224–31. doi:10.1002/1098-1004(200009)16:3<224::AID-HUMU5>3.0.CO;2-P. PMID 10980529. S2CID 3169748.
- ^ a b c d e f Online Mendelian Inheritance in Man, OMIM®. Johns Hopkins University, Baltimore, MD. MIM Number: {138140}: {08/21/2017}: . World Wide Web URL: https://omim.org/
- ^ Nelson DL, Cox MM (2008). Lehninger, Principles of Biochemistry. W. H. Freeman. ISBN 978-0-7167-7108-1.
- ^ Montel-Hagen A, Kinet S, Manel N, Mongellaz C, Prohaska R, Battini JL, Delaunay J, Sitbon M, Taylor N (March 2008). "Erythrocyte Glut1 triggers dehydroascorbic acid uptake in mammals unable to synthesize vitamin C". Cell. 132 (6): 1039–48. doi:10.1016/j.cell.2008.01.042. PMID 18358815. S2CID 18128118.*Lay summary in: "How Humans Make Up For An 'Inborn' Vitamin C Deficiency". ScienceDaily. March 21, 2008.
- ^ Uldry M, Thorens B (February 2004). "The SLC2 family of facilitated hexose and polyol transporters" (PDF). Pflügers Archiv. 447 (5): 480–9. doi:10.1007/s00424-003-1085-0. PMID 12750891. S2CID 25539725.
- ^ Seidner G, Alvarez MG, Yeh JI, O'Driscoll KR, Klepper J, Stump TS, Wang D, Spinner NB, Birnbaum MJ, De Vivo DC (February 1998). "GLUT-1 deficiency syndrome caused by haploinsufficiency of the blood–brain barrier hexose carrier". Nature Genetics. 18 (2): 188–91. doi:10.1038/ng0298-188. PMID 9462754. S2CID 7378231.
- ^ Manel N, Kim FJ, Kinet S, Taylor N, Sitbon M, Battini JL (November 2003). "The ubiquitous glucose transporter GLUT-1 is a receptor for HTLV". Cell. 115 (4): 449–59. doi:10.1016/S0092-8674(03)00881-X. PMID 14622599. S2CID 14399680.
- ^ North PE, Waner M, Mizeracki A, Mihm MC (January 2000). "GLUT1: a newly discovered immunohistochemical marker for juvenile hemangiomas". Human Pathology. 31 (1): 11–22. doi:10.1016/S0046-8177(00)80192-6. PMID 10665907.
- ^ Bunn RC, Jensen MA, Reed BC (April 1999). "Protein interactions with the glucose transporter binding protein GLUT1CBP that provide a link between GLUT1 and the cytoskeleton". Molecular Biology of the Cell. 10 (4): 819–32. doi:10.1091/mbc.10.4.819. PMC 25204. PMID 10198040.
- ^ Khan AA, Hanada T, Mohseni M, Jeong JJ, Zeng L, Gaetani M, Li D, Reed BC, Speicher DW, Chishti AH (May 2008). "Dematin and adducin provide a novel link between the spectrin cytoskeleton and human erythrocyte membrane by directly interacting with glucose transporter-1". The Journal of Biological Chemistry. 283 (21): 14600–9. doi:10.1074/jbc.M707818200. PMC 2386908. PMID 18347014.
- ^ Steinberg F, Gallon M, Winfield M, Thomas EC, Bell AJ, Heesom KJ, Tavaré JM, Cullen PJ (May 2013). "A global analysis of SNX27-retromer assembly and cargo specificity reveals a function in glucose and metal ion transport". Nature Cell Biology. 15 (5): 461–71. doi:10.1038/ncb2721. PMC 4052425. PMID 23563491.
- ^ Rungaldier S, Oberwagner W, Salzer U, Csaszar E, Prohaska R (March 2013). "Stomatin interacts with GLUT1/SLC2A1, band 3/SLC4A1, and aquaporin-1 in human erythrocyte membrane domains". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1828 (3): 956–66. doi:10.1016/j.bbamem.2012.11.030. PMC 3790964. PMID 23219802.
- ^ Lopez-Serra P, Marcilla M, Villanueva A, Ramos-Fernandez A, Palau A, Leal L, Wahi JE, Setien-Baranda F, Szczesna K, Moutinho C, Martinez-Cardus A, Heyn H, Sandoval J, Puertas S, Vidal A, Sanjuan X, Martinez-Balibrea E, Viñals F, Perales JC, Bramsem JB, Ørntoft TF, Andersen CL, Tabernero J, McDermott U, Boxer MB, Vander Heiden MG, Albar JP, Esteller M (April 2014). "A DERL3-associated defect in the degradation of SLC2A1 mediates the Warburg effect". Nature Communications. 5 (1): 3608. Bibcode:2014NatCo...5.3608L. doi:10.1038/ncomms4608. PMC 3988805. PMID 24699711.
- ^ Wood TE, Dalili S, Simpson CD, Hurren R, Mao X, Saiz FS, Gronda M, Eberhard Y, Minden MD, Bilan PJ, Klip A, Batey RA, Schimmer AD (November 2008). "A novel inhibitor of glucose uptake sensitizes cells to FAS-induced cell death". Molecular Cancer Therapeutics. 7 (11): 3546–55. doi:10.1158/1535-7163.MCT-08-0569. PMID 19001437. S2CID 7706108.
- ^ Siebeneicher H, Cleve A, Rehwinkel H, Neuhaus R, Heisler I, Müller T, Bauser M, Buchmann B (October 2016). "Identification and Optimization of the First Highly Selective GLUT1 Inhibitor BAY-876". ChemMedChem. 7 (11): 3546–55. doi:10.1002/cmdc.201600276. PMC 5095872. PMID 27552707.
Further reading
[edit]- Lankford J, Butler IJ, Koenig MK (June 2012). "Glucose transporter type I deficiency causing mitochondrial dysfunction". Journal of Child Neurology. 27 (6): 796–8. doi:10.1177/0883073811426503. PMID 22156785. S2CID 206549634.
- North PE, Waner M, Mizeracki A, Mihm MC (January 2000). "GLUT1: a newly discovered immunohistochemical marker for juvenile hemangiomas". Human Pathology. 31 (1): 11–22. doi:10.1016/S0046-8177(00)80192-6. PMID 10665907.
- Hruz PW, Mueckler MM (2001). "Structural analysis of the GLUT1 facilitative glucose transporter (review)". Molecular Membrane Biology. 18 (3): 183–93. doi:10.1080/09687680110072140. PMID 11681785. S2CID 218897534.
- Baumann MU, Deborde S, Illsley NP (October 2002). "Placental glucose transfer and fetal growth". Endocrine. 19 (1): 13–22. doi:10.1385/ENDO:19:1:13. PMID 12583599. S2CID 26301249.
- Mobasheri A, Richardson S, Mobasheri R, Shakibaei M, Hoyland JA (October 2005). "Hypoxia inducible factor-1 and facilitative glucose transporters GLUT1 and GLUT3: putative molecular components of the oxygen and glucose sensing apparatus in articular chondrocytes". Histology and Histopathology. 20 (4): 1327–38. doi:10.14670/HH-20.1327. PMID 16136514.
External links
[edit]- GeneReviews/NIH/UW entry on Glucose Transporter Type 1 Deficiency Syndrome
- Glucose+Transporter+Type+1 at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Overview of all the structural information available in the PDB for UniProt: P11166 (Solute carrier family 2, facilitated glucose transporter member 1) at the PDBe-KB.
This article incorporates text from the United States National Library of Medicine, which is in the public domain.