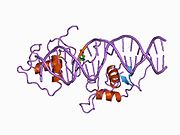
Rev-ErbA beta
 From Wikipedia the free encyclopedia
From Wikipedia the free encyclopedia
Rev-Erb beta (Rev-Erbβ), also known as nuclear receptor subfamily 1 group D member 2 (NR1D2), is a member of the Rev-Erb protein family. Rev-Erbβ, like Rev-Erbα, belongs to the nuclear receptor superfamily of transcription factors and can modulate gene expression through binding to gene promoters.[5] Together with Rev-Erbα, Rev-Erbβ functions as a major regulator of the circadian clock. These two proteins are partially redundant.[6] Current research suggests that Rev-Erbβ is less important in maintaining the circadian clock than Rev-Erbα; knock-out studies of Rev-Erbα result in significant circadian disruption but the same has not been found with Rev-Erbβ. Rev-Erbβ compensation for Rev-Erbα varies across tissues, and further research is needed to elucidate the separate role of Rev-Erbβ.[7]
This gene is expressed in the central and peripheral nervous system, spleen, mandibular maxillary processes, and blood islands. Rev-Erbβ plays a major role in the conduction of inductive signals to aid in controlling differentiating neurons.[5]
Discovery
[edit]Rev-Erbβ was discovered in 1994, when B. Dumas et al. isolated its cDNA, naming the new receptor BD73.[5] The name Rev-Erbβ was coined a few months later in a paper by Eva Enmark, Tommi Kainu, Markku Tapio Pelto-Huikko, and Jan Ǻke Gustafsson where they isolated Rev-Erb alpha cDNA in a rat brain.[8]
A new isoform of Rev-Erbβ, named Rev-Erbβ 2, was discovered using rat cDNA a few months later in 1995 by N. Giambiagi and colleagues.[7] They found it to be identical to Rev-Erbβ 1, except that the Rev-Erbβ 1 protein is 195 amino acids longer than Rev-Erbβ 2. However, further research has indicated that the discovered Rev-Erbβ 2 cDNA was likely a splice variant of the Nr1d2 gene that arose through alternative splicing and the use of a different polyadenylation site.
Genetics and Evolution
[edit]In mammals, the NR1D2 (nuclear receptor subfamily 1 group D member 2) gene encodes the protein Rev-Erbβ. Unlike NR1D1, the strand opposite NR1D2 does not have any significant reading frames, and the gene is located on the forward strand of chromosome 3.[9] Despite their different locations, the NR1D1 and NR1D2 genes are highly homologous and are paralogs within the genome.[5] In humans, the NR1D2 gene itself contains 10 exons which form 5 splice variants (NR1D2-201 - NR1D2-205), ranging from 5231 base pairs (NR1D2-201) to 600 base pairs (NR1D2-204). However, only NR1D2-201 produces a functional protein. In mammals, NR1D2 (Rev-Erbβ) is expressed throughout the body and with high expression in several tissues, including the brain, liver, skeletal muscle, and adipose tissue.[9]
Comparison of the human NR1D2 sequence with other species indicates a high level of conservation across animals, with 472 discovered orthologs, including in mice, chickens, lizards, and zebrafish. Similarly to NR1D1, this suggests NR1D2 was present in the most recent common animal ancestor. NR1D2 has only one paralog in humans, the NR1D1 gene, which is located on chromosome 17, but it is closely related to other members of the nuclear receptor family and is functionally related to other nuclear receptor genes, such as thyroid hormone receptor beta (THRB), peroxisome proliferator activated receptor delta (PPARD), and retinoic acid receptor beta (RARB). Linkage analysis reveals that NR1D2 and THRB are highly linked due to proximity on chromosome 3, and that they are both linked to RARB. Combined with the linkage between the NR1D1/THRA locus and the RARA gene, this suggests that these two gene clusters arose from a duplication event.[10]
Structure
[edit]
The human NR1D2 gene produces a protein product (REV-ERBβ) of 579 amino acids. Rev-Erbβ is similar to Rev-Erbα in both its structure and mechanism of transcriptional repression. Like Rev-Erbα, Rev-Erbβ has 3 major functional domains which are common to nuclear receptor proteins, including a DNA-binding domain (DBD) and a ligand-binding domain (LBD) at the C-terminus, which are highly conserved in Rev-Erb orthologs, and a N-terminus domain which allows for activity modulation.[11]
Much like Rev-Erbα, Rev-Erbβ can bind to two classes of DNA response elements via its DBD, which contains two C4-type zinc fingers.[12] These two classes include a DNA sequence commonly referred to as RORE due to its interaction with the transcriptional activator Retinoic Acid Receptor-related Orphan Receptor (ROR) and a direct repeat 2 element of RORE known as RevDR2.[13] The Rev-Erb proteins are unique from other nuclear receptors in that they do not have a helix in the C-terminal that is necessary for coactivator recruitment and activation by nuclear receptors via their LBD. Instead, the Rev-Erbs can repress transcription as a monomer through competitive binding at single RORE elements by preventing the binding of constitutive transcription activator ROR or as a homodimer through binding to RevDR2 sites.[14] The Rev-Erb homodimer is required for its interaction with Nuclear Receptor Co-Repressor (NCoR), or more weakly, with Silencing Mediator of Retinoid and Thyroid Receptors (SMRT). The interaction with NCoR is stabilized by interaction with heme, which binds the [clarification needed] to the Rev-Erb ligand-binding pocket. Rev-Erbβ undergoes a conformational change when complexed with heme, as its structure shows that helices 3,7, and 11 move to enlarge the ligand binding pocket in order to accommodate heme. The repression by Rev-Erb proteins also requires interaction of class I histone deacetylase 3 (HDAC3) with NCoR, which results in gene repression via histone deacetylation.[12]
Function
[edit]Circadian oscillator
[edit]Rev-Erbβ binds to genomic Rev-Erbα-binding sites that have a diurnal profile identical or similar to Rev-Erbα. This protein also helps maintain clock and metabolic gene regulation and protects system functioning when Rev-Erbα is missing. Rev-Erbβ compensates for loss of function from metabolic distress in the case that Rev-Erbα is lost. The liver and metabolic processes can still run when Rev-Erbα is missing and Rev-Erbβ is present. Losing both Rev-Erbα and Rev-Erbβ causes cells to become arrhythmic.[15]
When Rev-Erbβ is missing, there can be significant change in performance of metabolic activity with drastic effects. For example:
- Rev-Erbβ deficiency causes a drastic difference in the coupled formation of circadian networks of gene expression, while core clock gene expression remains oscillating.
- Having neither Rev-Erbα or Rev-Erbβ does not affect expression rhythms of core clock genes but affects other rhythmically-expressed output genes.
- Rev-Erbβ deficiency does not change circadian expression rhythms of PER2.[15]
Metabolism
[edit]Rev-Erbβ plays a role in blocking the trans-activation of retinoic acid-related orphan receptor-α (RORα). RORα is involved in the regulation of lipoprotein cholesterol, lipid homeostasis, and inflammation. Rev-Erbβ and RORα are both expressed in similar tissues, such as skeletal muscle. They have similar expression patterns, target genes, and cognate sequences within the skeletal muscle. Rev-Erbβ causes several genes assisting in lipid absorption to decrease expression. Rev-Erbβ controls lipid and energy homoeostasis in skeletal muscle. Rev-Erbβ may be useful in therapeutic treatments of dyslipidemia and regulating muscle growth.[16]
Rev-Erbβ is also a circadian regulated gene; its mRNA displays rhythmic expression in vivo and in serum-synchronized cell cultures. However, it is currently unknown to what extent Rev-Erbβ contributes to oscillations of the core circadian clock. However it has been shown that heme suppresses hepatic gluconeogenic gene expression and glucose output through the related Rev-Erbα receptor which mediates gene repression. Hence, the Rev-Erbα receptor detects heme and thereby coordinates the cellular clock, glucose homeostasis, and energy metabolism.[17]
Rev-Erbβ plays a role in skeletal muscle mitochondrial biogenesis. Originally Rev-Erbβ was thought to be functionally redundant of Rev-Erbα but recent findings prove that there are subtle differences. Rev-Erbβ ligands may be used in the treatment of metabolic disorders, like metabolic syndrome. It has control of skeletal muscle metabolism and energy that can be beneficial in treatment options.[18]
Rev-Erbβ gene contributes to the downstream regulation of clock output genes by generating specific KO mutants. It is still unknown all of the functions Rev-Erbβ has in the core circadian clock and exactly how it differs from Rev-Erbα.
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000174738 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000021775 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c d Dumas B, Harding HP, Choi HS, Lehmann KA, Chung M, Lazar MA, Moore DD (August 1994). "A new orphan member of the nuclear hormone receptor superfamily closely related to Rev-Erb". Molecular Endocrinology. 8 (8): 996–1005. doi:10.1210/mend.8.8.7997240. PMID 7997240.
- ^ Koh YS, Moore DD (April 1999). "Linkage of the nuclear hormone receptor genes NR1D2, THRB, and RARB: evidence for an ancient, large-scale duplication". Genomics. 57 (2): 289–92. doi:10.1006/geno.1998.5683. PMID 10198169.
- ^ a b Giambiagi N, Cassia R, Petropoulos I, Part D, Cereghini S, Zakin MM, Ochoa A (December 1995). "Rev-erb beta 2, a novel isoform of the Rev-erb family of orphan nuclear receptors". Biochemistry and Molecular Biology International. 37 (6): 1091–1102. PMID 8747539.
- ^ Enmark E, Kainu T, Pelto-Huikko M, Gustafsson JA (October 1994). "Identification of a novel member of the nuclear receptor superfamily which is closely related to Rev-ErbA". Biochemical and Biophysical Research Communications. 204 (1): 49–56. doi:10.1006/bbrc.1994.2424. PMID 7945391.
- ^ a b Yates AD, Achuthan P, Akanni W, Allen J, Allen J, Alvarez-Jarreta J, et al. (January 2020). "Ensembl 2020". Nucleic Acids Research. 48 (D1): D682–D688. doi:10.1093/nar/gkz966. PMC 7145704. PMID 31691826.
- ^ Burris TP (July 2008). "Nuclear hormone receptors for heme: REV-ERBalpha and REV-ERBbeta are ligand-regulated components of the mammalian clock". Molecular Endocrinology. 22 (7): 1509–20. doi:10.1210/me.2007-0519. PMC 5419435. PMID 18218725.
- ^ Woo EJ, Jeong DG, Lim MY, Jun Kim S, Kim KJ, Yoon SM, et al. (October 2007). "Structural insight into the constitutive repression function of the nuclear receptor Rev-erbbeta". Journal of Molecular Biology. 373 (3): 735–44. doi:10.1016/j.jmb.2007.08.037. PMID 17870090.
- ^ a b Pardee KI, Xu X, Reinking J, Schuetz A, Dong A, Liu S, et al. (February 2009). "The structural basis of gas-responsive transcription by the human nuclear hormone receptor REV-ERBbeta". PLOS Biology. 7 (2): e43. doi:10.1371/journal.pbio.1000043. PMC 2652392. PMID 19243223.
- ^ "NR1D1 Gene | NR1D1 Protein | NR1D1 Antibody". GeneCards. Retrieved 2021-05-06.
- ^ Zhao Q, Khorasanizadeh S, Miyoshi Y, Lazar MA, Rastinejad F (May 1998). "Structural elements of an orphan nuclear receptor-DNA complex". Molecular Cell. 1 (6): 849–61. doi:10.1016/s1097-2765(00)80084-2. PMID 9660968.
- ^ a b Bugge A, Feng D, Everett LJ, Briggs ER, Mullican SE, Wang F, et al. (April 2012). "Rev-erbα and Rev-erbβ coordinately protect the circadian clock and normal metabolic function". Genes & Development. 26 (7): 657–67. doi:10.1101/gad.186858.112. PMC 3323877. PMID 22474260.
- ^ a b Ikeda R, Tsuchiya Y, Koike N, Umemura Y, Inokawa H, Ono R, et al. (July 2019). "REV-ERBα and REV-ERBβ function as key factors regulating Mammalian Circadian Output". Scientific Reports. 9 (1): 10171. Bibcode:2019NatSR...910171I. doi:10.1038/s41598-019-46656-0. PMC 6629614. PMID 31308426.
- ^ Ramakrishnan SN, Lau P, Burke LJ, Muscat GE (March 2005). "Rev-erbbeta regulates the expression of genes involved in lipid absorption in skeletal muscle cells: evidence for cross-talk between orphan nuclear receptors and myokines". The Journal of Biological Chemistry. 280 (10): 8651–9. doi:10.1074/jbc.M413949200. PMID 15623503.
- ^ Yin L, Wu N, Curtin JC, Qatanani M, Szwergold NR, Reid RA, et al. (December 2007). "Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways". Science. 318 (5857): 1786–9. Bibcode:2007Sci...318.1786Y. doi:10.1126/science.1150179. PMID 18006707. S2CID 84073753.
Further reading
[edit]- Ramakrishnan SN, Lau P, Crowther LM, Cleasby ME, Millard S, Leong GM, Cooney GJ, Muscat GE (October 2009). "Rev-erb beta regulates the Srebp-1c promoter and mRNA expression in skeletal muscle cells" (PDF). Biochemical and Biophysical Research Communications. 388 (4): 654–9. doi:10.1016/j.bbrc.2009.08.045. PMID 19682428.
External links
[edit]- NR1D2+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)